Which Of The Following Unwinds The Helix To Provide Single Stranded Template
Affiliate 9: Introduction to Molecular Biology
9.2 Deoxyribonucleic acid Replication
Learning Objectives
Past the end of this section, you will exist able to:
- Explicate the process of DNA replication
- Explain the importance of telomerase to DNA replication
- Describe mechanisms of DNA repair
When a cell divides, it is important that each daughter jail cell receives an identical copy of the Dna. This is accomplished past the process of Deoxyribonucleic acid replication. The replication of DNA occurs during the synthesis phase, or S stage, of the cell wheel, before the cell enters mitosis or meiosis.
The elucidation of the structure of the double helix provided a hint as to how Deoxyribonucleic acid is copied. Recall that adenine nucleotides pair with thymine nucleotides, and cytosine with guanine. This means that the two strands are complementary to each other. For example, a strand of Dna with a nucleotide sequence of AGTCATGA volition have a complementary strand with the sequence TCAGTACT (Figure ix.8).
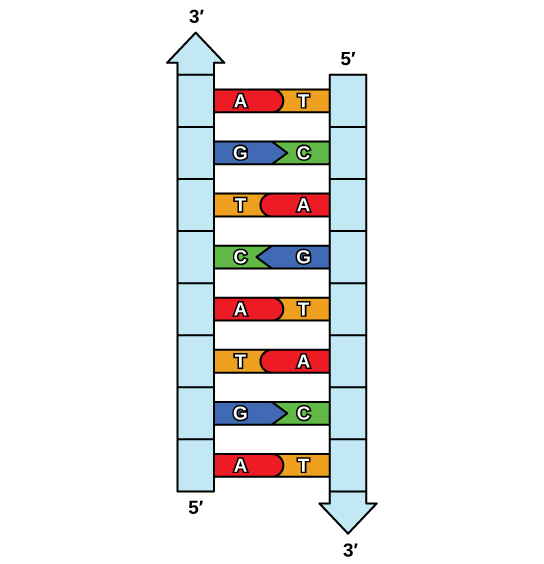
Because of the complementarity of the two strands, having ane strand ways that it is possible to recreate the other strand. This model for replication suggests that the two strands of the double helix split up during replication, and each strand serves as a template from which the new complementary strand is copied (Effigy 9.9).
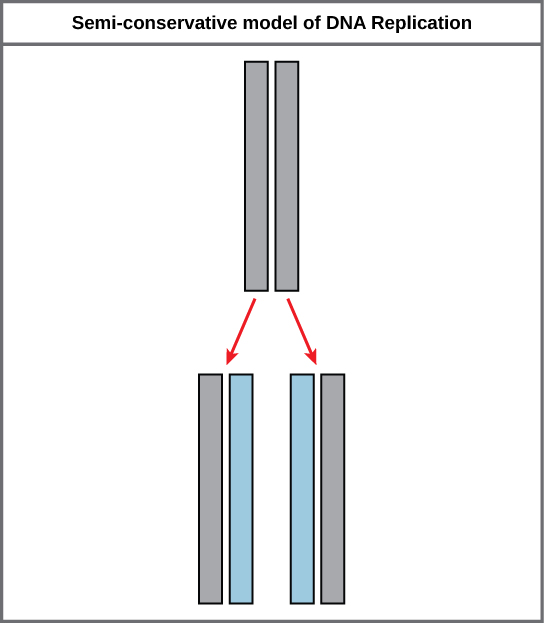
During DNA replication, each of the two strands that brand up the double helix serves equally a template from which new strands are copied. The new strand will be complementary to the parental or "former" strand. Each new double strand consists of one parental strand and 1 new daughter strand. This is known equally semiconservative replication. When two Dna copies are formed, they have an identical sequence of nucleotide bases and are divided equally into two daughter cells.
Dna Replication in Eukaryotes
Because eukaryotic genomes are very complex, Deoxyribonucleic acid replication is a very complicated procedure that involves several enzymes and other proteins. It occurs in three chief stages: initiation, elongation, and termination.
Recall that eukaryotic Dna is spring to proteins known every bit histones to grade structures called nucleosomes. During initiation, the DNA is made accessible to the proteins and enzymes involved in the replication procedure. How does the replication machinery know where on the Dna double helix to brainstorm? It turns out that there are specific nucleotide sequences called origins of replication at which replication begins. Certain proteins bind to the origin of replication while an enzyme chosen helicase unwinds and opens up the DNA helix. Equally the Deoxyribonucleic acid opens upwardly, Y-shaped structures called replication forks are formed (Figure 9.10). Two replication forks are formed at the origin of replication, and these go extended in both directions equally replication proceeds. There are multiple origins of replication on the eukaryotic chromosome, such that replication can occur simultaneously from several places in the genome.
During elongation, an enzyme called Dna polymerase adds DNA nucleotides to the 3′ finish of the template. Because DNA polymerase can only add new nucleotides at the end of a backbone, a primer sequence, which provides this starting point, is added with complementary RNA nucleotides. This primer is removed afterward, and the nucleotides are replaced with Dna nucleotides. One strand, which is complementary to the parental Deoxyribonucleic acid strand, is synthesized continuously toward the replication fork so the polymerase tin can add nucleotides in this direction. This continuously synthesized strand is known as the leading strand. Considering Dna polymerase tin only synthesize DNA in a 5′ to iii′ management, the other new strand is put together in short pieces chosen Okazaki fragments. The Okazaki fragments each require a primer made of RNA to start the synthesis. The strand with the Okazaki fragments is known as the lagging strand. As synthesis proceeds, an enzyme removes the RNA primer, which is and so replaced with Dna nucleotides, and the gaps betwixt fragments are sealed past an enzyme chosen Dna ligase.
The process of Deoxyribonucleic acid replication tin be summarized as follows:
- Deoxyribonucleic acid unwinds at the origin of replication.
- New bases are added to the complementary parental strands. One new strand is made continuously, while the other strand is made in pieces.
- Primers are removed, new Dna nucleotides are put in identify of the primers and the backbone is sealed past DNA ligase.
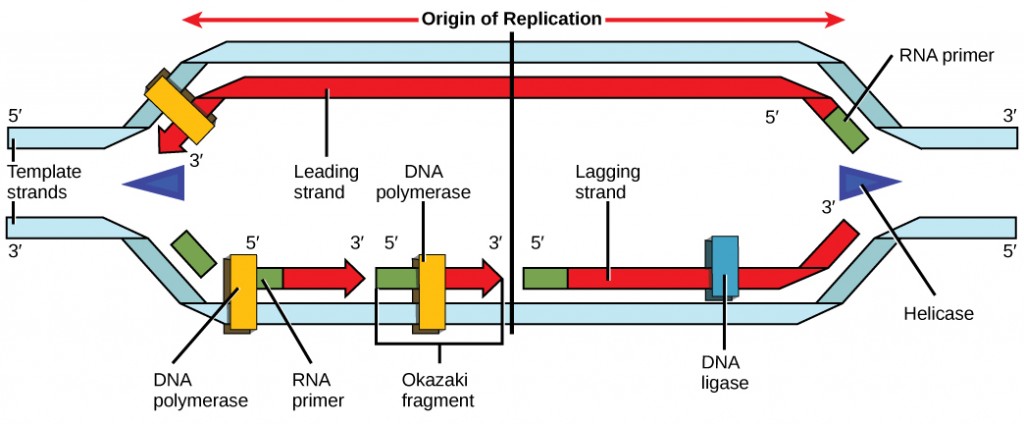
Yous isolate a prison cell strain in which the joining together of Okazaki fragments is impaired and suspect that a mutation has occurred in an enzyme institute at the replication fork. Which enzyme is well-nigh likely to be mutated?
Telomere Replication
Considering eukaryotic chromosomes are linear, DNA replication comes to the finish of a line in eukaryotic chromosomes. As you take learned, the Deoxyribonucleic acid polymerase enzyme can add nucleotides in merely one direction. In the leading strand, synthesis continues until the cease of the chromosome is reached; still, on the lagging strand at that place is no place for a primer to be made for the Deoxyribonucleic acid fragment to be copied at the end of the chromosome. This presents a problem for the jail cell considering the ends remain unpaired, and over fourth dimension these ends get progressively shorter as cells proceed to divide. The ends of the linear chromosomes are known as telomeres, which accept repetitive sequences that do not code for a particular cistron. Equally a consequence, it is telomeres that are shortened with each round of DNA replication instead of genes. For instance, in humans, a six base-pair sequence, TTAGGG, is repeated 100 to chiliad times. The discovery of the enzyme telomerase (Figure 9.11) helped in the understanding of how chromosome ends are maintained. The telomerase attaches to the end of the chromosome, and complementary bases to the RNA template are added on the end of the DNA strand. One time the lagging strand template is sufficiently elongated, DNA polymerase can now add nucleotides that are complementary to the ends of the chromosomes. Thus, the ends of the chromosomes are replicated.
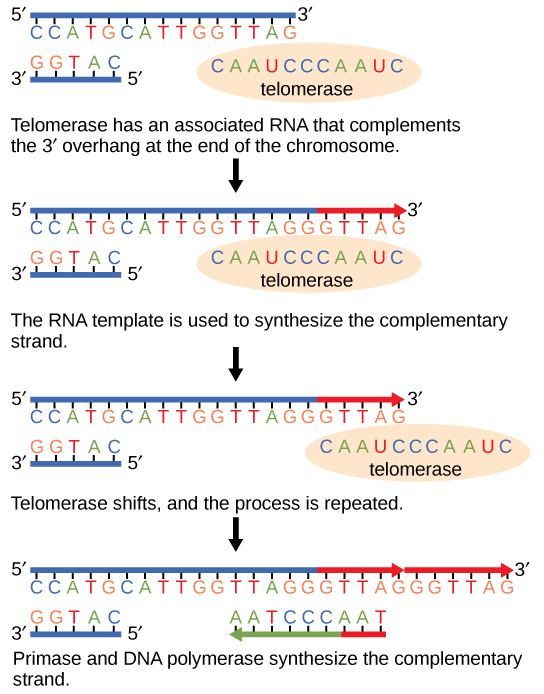
Telomerase is typically found to exist active in germ cells, adult stem cells, and some cancer cells. For her discovery of telomerase and its action, Elizabeth Blackburn (Figure 9.12) received the Nobel Prize for Medicine and Physiology in 2009.

Telomerase is non active in adult somatic cells. Adult somatic cells that undergo jail cell division proceed to have their telomeres shortened. This essentially ways that telomere shortening is associated with aging. In 2010, scientists found that telomerase can opposite some age-related conditions in mice, and this may have potential in regenerative medicine. one Telomerase-deficient mice were used in these studies; these mice take tissue atrophy, stem-cell depletion, organ organisation failure, and dumb tissue injury responses. Telomerase reactivation in these mice acquired extension of telomeres, reduced Dna damage, reversed neurodegeneration, and improved functioning of the testes, spleen, and intestines. Thus, telomere reactivation may have potential for treating age-related diseases in humans.
DNA Replication in Prokaryotes
Call up that the prokaryotic chromosome is a circular molecule with a less extensive coiling structure than eukaryotic chromosomes. The eukaryotic chromosome is linear and highly coiled effectually proteins. While there are many similarities in the Deoxyribonucleic acid replication process, these structural differences necessitate some differences in the DNA replication process in these ii life forms.
DNA replication has been extremely well-studied in prokaryotes, primarily because of the small size of the genome and large number of variants bachelor. Escherichia coli has 4.6 1000000 base pairs in a single circular chromosome, and all of information technology gets replicated in approximately 42 minutes, starting from a single origin of replication and proceeding around the chromosome in both directions. This ways that approximately g nucleotides are added per second. The process is much more rapid than in eukaryotes. The table below summarizes the differences betwixt prokaryotic and eukaryotic replications.
| Property | Prokaryotes | Eukaryotes |
|---|---|---|
| Origin of replication | Single | Multiple |
| Charge per unit of replication | m nucleotides/s | fifty to 100 nucleotides/southward |
| Chromosome construction | round | linear |
| Telomerase | Not present | Present |
Concept in Activeness

Click through a tutorial on Dna replication.
DNA Repair
Deoxyribonucleic acid polymerase can make mistakes while adding nucleotides. It edits the Deoxyribonucleic acid by proofreading every newly added base. Incorrect bases are removed and replaced by the right base of operations, and then polymerization continues (Figure ix.13 a). Most mistakes are corrected during replication, although when this does not happen, the mismatch repair mechanism is employed. Mismatch repair enzymes recognize the wrongly incorporated base and excise information technology from the Deoxyribonucleic acid, replacing information technology with the correct base (Figure 9.13 b). In however some other blazon of repair, nucleotide excision repair, the Deoxyribonucleic acid double strand is unwound and separated, the incorrect bases are removed forth with a few bases on the 5′ and three′ end, and these are replaced by copying the template with the assistance of Deoxyribonucleic acid polymerase (Effigy 9.13 c). Nucleotide excision repair is particularly important in correcting thymine dimers, which are primarily acquired by ultraviolet light. In a thymine dimer, two thymine nucleotides adjacent to each other on one strand are covalently bonded to each other rather than their complementary bases. If the dimer is not removed and repaired information technology will lead to a mutation. Individuals with flaws in their nucleotide excision repair genes show extreme sensitivity to sunlight and develop pare cancers early in life.
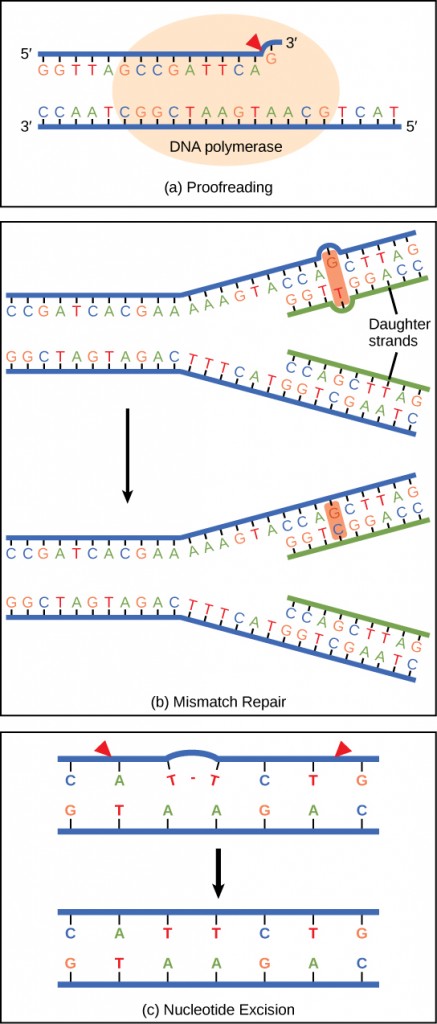
Virtually mistakes are corrected; if they are not, they may result in a mutation—defined as a permanent change in the Dna sequence. Mutations in repair genes may lead to serious consequences like cancer.
Section Summary
Deoxyribonucleic acid replicates by a semi-conservative method in which each of the two parental Deoxyribonucleic acid strands act every bit a template for new Deoxyribonucleic acid to be synthesized. Subsequently replication, each DNA has 1 parental or "onetime" strand, and one girl or "new" strand.
Replication in eukaryotes starts at multiple origins of replication, while replication in prokaryotes starts from a single origin of replication. The DNA is opened with enzymes, resulting in the formation of the replication fork. Primase synthesizes an RNA primer to initiate synthesis by DNA polymerase, which tin can add nucleotides in just one management. Ane strand is synthesized continuously in the direction of the replication fork; this is called the leading strand. The other strand is synthesized in a direction away from the replication fork, in short stretches of Dna known as Okazaki fragments. This strand is known as the lagging strand. Once replication is completed, the RNA primers are replaced by DNA nucleotides and the Dna is sealed with DNA ligase.
The ends of eukaryotic chromosomes pose a problem, as polymerase is unable to extend them without a primer. Telomerase, an enzyme with an inbuilt RNA template, extends the ends by copying the RNA template and extending ane end of the chromosome. DNA polymerase tin can then extend the Dna using the primer. In this manner, the ends of the chromosomes are protected. Cells have mechanisms for repairing DNA when it becomes damaged or errors are fabricated in replication. These mechanisms include mismatch repair to replace nucleotides that are paired with a not-complementary base and nucleotide excision repair, which removes bases that are damaged such as thymine dimers.
Glossary
DNA ligase: the enzyme that catalyzes the joining of DNA fragments together
Deoxyribonucleic acid polymerase: an enzyme that synthesizes a new strand of DNA complementary to a template strand
helicase: an enzyme that helps to open up upwardly the DNA helix during Dna replication past breaking the hydrogen bonds
lagging strand: during replication of the 3′ to 5′ strand, the strand that is replicated in curt fragments and away from the replication fork
leading strand: the strand that is synthesized continuously in the 5′ to iii′ direction that is synthesized in the direction of the replication fork
mismatch repair: a form of DNA repair in which not-complementary nucleotides are recognized, excised, and replaced with correct nucleotides
mutation: a permanent variation in the nucleotide sequence of a genome
nucleotide excision repair: a grade of Dna repair in which the DNA molecule is unwound and separated in the region of the nucleotide damage, the damaged nucleotides are removed and replaced with new nucleotides using the complementary strand, and the Deoxyribonucleic acid strand is resealed and allowed to rejoin its complement
Okazaki fragments: the Deoxyribonucleic acid fragments that are synthesized in short stretches on the lagging strand
primer: a short stretch of RNA nucleotides that is required to initiate replication and allow DNA polymerase to bind and begin replication
replication fork: the Y-shaped construction formed during the initiation of replication
semiconservative replication: the method used to replicate DNA in which the double-stranded molecule is separated and each strand acts as a template for a new strand to be synthesized, so the resulting DNA molecules are equanimous of 1 new strand of nucleotides and one onetime strand of nucleotides
telomerase: an enzyme that contains a catalytic part and an inbuilt RNA template; it functions to maintain telomeres at chromosome ends
telomere: the Deoxyribonucleic acid at the end of linear chromosomes
Footnotes
1 Mariella Jaskelioff, et al., "Telomerase reactivation reverses tissue degeneration in aged telomerase-deficient mice," Nature, 469 (2011):102–seven.
Which Of The Following Unwinds The Helix To Provide Single Stranded Template,
Source: https://opentextbc.ca/biology/chapter/9-2-dna-replication/
Posted by: wilsonsirtho.blogspot.com


0 Response to "Which Of The Following Unwinds The Helix To Provide Single Stranded Template"
Post a Comment